DeepMind’s latest AlphaFold model is more useful for drug discovery

Nearly five years ago, DeepMind, one of Google’s more prolific AI-centered research labs, debuted AlphaFold, an AI system that can accurately predict the structures of many proteins inside the human body. Since then, DeepMind has improved on the system, releasing an updated and more capable version of AlphaFold — AlphaFold 2 — in 2020.
And the lab’s work continues.
Today, DeepMind revealed that the newest release of AlphaFold, the successor to AlphaFold 2, can generate predictions for nearly all molecules in the Protein Data Bank, the world’s largest open access database of biological molecules.
Already, Isomorphic Labs, a spin-off of DeepMind focused on drug discovery, is applying the new AlphaFold model — which it co-designed — to therapeutic drug design, according to a post on the DeepMind blog, helping to characterize different types of molecular structures important for treating disease.
New capabilities
The new AlphaFold’s capabilities extend beyond protein prediction.
DeepMind claims that the model can also accurately predict the structures of ligands — molecules that bind to “receptor” proteins and cause changes in how cells communicate — as well as nucleic acids (molecules that contain key genetic information) and post-translational modifications (chemical changes that occur after a protein’s created).
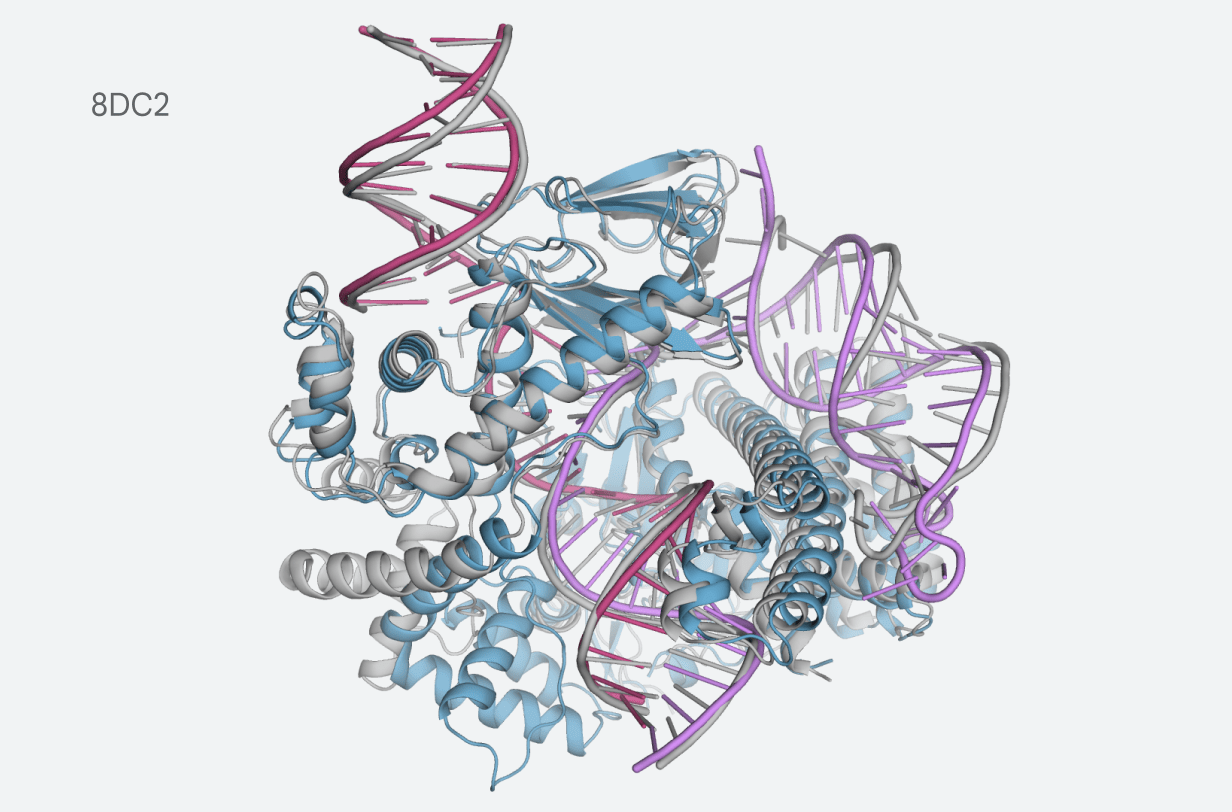
Image Credits: DeepMind
Predicting protein-ligand structures can be a useful tool in drug discovery, DeepMind notes, as it can help scientists identify and design new molecules that could become drugs.
Currently, pharmaceutical researchers use computer simulations known as “docking methods” to determine how proteins and ligands will interact. Docking methods require specifying a reference protein structure and a suggested position on that structure for the ligand to bind to.
With the latest AlphaFold, however, there’s no need to use a reference protein structure or suggested position. The model can predict proteins that haven’t been “structurally characterized” before, while at the same time simulating how proteins and nucleic acids interact with other molecules — a level of modeling that DeepMind says isn’t possible with today’s docking methods.
“Early analysis also shows that our model greatly outperforms [the previous generation of] AlphaFold on some protein structure prediction problems that are relevant for drug discovery, like antibody binding,” DeepMind writes in the post. “Our model’s dramatic leap in performance shows the potential of AI to greatly enhance scientific understanding of the molecular machines that make up the human body.”
The newest AlphaFold isn’t perfect, though.
In a whitepaper detailing the system’s strengths and limitations, researchers at DeepMind and Isomorphic Labs reveal that the system falls short of the best-in-class method for predicting the structures of RNA molecules — the molecules in the body that carry the instructions for making proteins.
Doubtless, both DeepMind and Isomorphic Labs are working to address this.


